A new study published on arXiv warns that more pre-training data may not always lead to better AI models. Researchers from prestigious institutions, including Carnegie Mellon University, Stanford University, Harvard University, and Princeton University, have highlighted a concerning phenomenon known as "Catastrophic Overtraining." Their research suggests that extending pre-training can actually degrade a model’s ability to be fine-tuned effectively, leading to poorer performance in real-world applications.
The conventional wisdom in the AI industry has been that more data equals better performance. However, this study challenges that assumption. The researchers stated, “Contrary to common belief, longer pre-training does not always lead to better post-trained models.” They argue that as models are pre-trained on more tokens, they become increasingly sensitive to perturbations, which can hinder their adaptability during fine-tuning.
To illustrate their findings, the researchers compared two versions of Ai2’s open-source OLMo-1B model. One version was trained on 2.3 trillion tokens, while the other was trained on a staggering 3 trillion tokens. Surprisingly, the model that was trained on more data performed worse after fine-tuning, showing a 2-3% lower accuracy on standard benchmarks such as ARC-Challenge, PIQA, and AlpacaEval.
The authors explain this degradation in performance through a concept they term “progressive sensitivity.” As models are trained for longer durations, their internal parameters become increasingly sensitive to changes, such as adjustments made during fine-tuning or the introduction of additional data. This heightened sensitivity means that even minor changes or small amounts of noise in the data can significantly disrupt what the model has already learned.
Further supporting their findings, the researchers conducted experiments where they added Gaussian noise to pre-trained models. They found that the performance of these models deteriorated significantly with an increase in pre-training tokens. Additionally, they validated their results using a different setup involving fine-tuned benchmarks, which yielded similar outcomes.
The researchers caution that their findings are not universally applicable, noting that the risk of catastrophic overtraining is particularly pronounced in smaller models. They emphasized, “Catastrophic overtraining may be inevitable, even if the fine-tuning process is regularized, especially when the pre-training and fine-tuning tasks are misaligned.” This highlights the critical importance of ensuring alignment between training and fine-tuning objectives.
As AI model pre-training remains a crucial component of the development process, the implications of this study are significant. The researchers suggest that developers may need to rethink their approach to building AI models. Instead of simply scaling up data and model size, they advocate for optimizing the entire training pipeline. “Our findings call for a renewed focus on model scaling that considers the entire training pipeline,” they concluded.
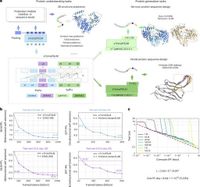
In parallel, another breakthrough in the field of AI has emerged with the introduction of a unified protein language model known as xTrimoPGLM. This model addresses both protein understanding and generation tasks through an innovative pretraining framework. Trained at an unprecedented scale of 100 billion parameters and 1 trillion training tokens, xTrimoPGLM has shown remarkable success in learning biological information from protein sequences.
The xTrimoPGLM model outperforms other advanced baselines across 18 protein understanding benchmarks spanning four categories. Additionally, it facilitates an atomic-resolution view of protein structures, leading to an advanced three-dimensional structural prediction model that surpasses existing language model-based tools.
Notably, xTrimoPGLM is capable of generating de novo protein sequences that adhere to the principles of natural proteins. Moreover, it can perform programmable generation after supervised fine-tuning on curated sequences. These capabilities underscore the model's substantial versatility in understanding and generating protein sequences, which is particularly relevant in the evolving landscape of foundation models in protein science.
The pretraining dataset for xTrimoPGLM was curated from two extensive data repositories: UniRef90 (version preceding December 2022) and ColabFoldDB. All structure prediction datasets utilized in the research are sourced from the AlphaFold database and the Protein Data Bank (PDB) from May 2020.
The training of xTrimoPGLM employed DeepSpeed v0.6.1, with data analysis conducted using Python v3.8, along with various libraries including NumPy, SciPy, Seaborn, Matplotlib, and Pandas. The researchers utilized TM-align v20190822 for computing TM-scores, and the structure visualizations were created in Pymol v2.3.0. Protein 3D structures were predicted using AlphaFold2 with the official implementations.
The composition of the dataset used for pre-training xTrimoPGLM is noteworthy. It includes sequences from four superkingdoms: Bacteria (67%), Archaea (3%), Eukarya (27%), and Viruses (1%), with 2% of sequences labeled as unclassified. The dataset also encompasses 17 classified kingdoms and a total of 273 known phyla.
As AI continues to evolve, the findings from both studies highlight the need for a nuanced understanding of model training processes. While the quest for larger datasets and more extensive pre-training remains a driving force in AI development, researchers are now advocating for a more balanced approach that considers the overall training strategy. By adopting smarter strategies for AI development, the notion that less can sometimes be more may well become a guiding principle in the future.